ABSTRACT
Background
This study focuses on the computational (in silico) analysis of 1H-phenothiazine derivatives to estimate their pharmacological potential, particularly as inhibitors of monoamine oxidase (PDB ID: 2BXR), an enzyme linked to depression.
Materials and Methds
Using assorted computational methods, the study investigates structure-activity relationships and predicts the impact of structural changes on drug effectiveness.
Results
Results show that several compounds demonstrated significant cytotoxic activity against depression-related targets, suggesting promising inhibitory and anti-inflammatory properties. The aim of this work was to examine the inflammatory and enzyme-inhibitory properties of a novel series of 1H-phenothiazin derivatives in order to determine whether they may be used as multi-action therapeutic drugs. Combi Lab studies and 3D-QSAR were carried out using the Molecular Design Suite. Molecular docking analysis was conducted using Schrodinger Maestro.
Conclusion
Out of the sixteen compounds made utilizing a Combinatorial technique, five compounds (PS6; PS5; PS11; PS13, and PS15) showed greater projected biological activity compared to the dataset’s most active molecule. The amino acid residues on monoamine oxidase (PDB ID: 2BXR) (PDB: 3LN1), including Tyr-385, Trp-387, Phe-518, Gly-526, Met-522, Tyr348, Val-349, Leu-352, Phe381, Leu-384, and Tyr-385were in close proximity to these substances.
INTRODUCTION
Current antidepressants usually work by affecting brain chemicals like serotonin and noradrenaline, which are essential for mood regulation. The most commonly used drugs include selective serotonin reuptake inhibitors such as fluoxetine, citalopram, escitalopram, fluvoxamine, as well as noradrenaline reuptake inhibitors like and desipramine. Drug discovery is a very complex and costly process that involves choosing a disease, identifying and confirming the right targets, finding and improving lead compounds, and then testing them through preclinical and clinical trials. Research has shown that many drug candidates fail because of poor pharmacokinetic profiles and ADMET properties. This has made it important to assess “drug-likeness” early on in the development process. Computer-based methods are now key in early drug discovery to help predict and reduce the risk of toxicity. Phenothiazine has been widely used in medicinal chemistry, and their derivatives are known for various activities, including antipsychotic and antidepressant effects. A popular method for forecasting how a tiny molecule would behave is called molecular docking (drug) fits into the binding site of a protein, offering insights into both the binding shape and strength. This method is very useful for both screening potential drug candidates and analyzing how well lead compounds might work. It might be difficult to treat multifactorial illnesses since they frequently have several underlying causes. There are a lot of promise for treating a wide range of illnesses by creating innovative hybrid molecules with a variety of pharmacological actions that can alter different biochemical pathways (Boehmet al., 1996). Despite the tremendous Designing and optimizing such compounds is challenging, compared to medicine combinations or multicomponent therapies, this method has clear advantages because it lowers the possibility of drug interactions. Atypical antipsychotics, such as olanzapine, risperidone and aripiprazole, predominantly function as antagonists at dopamine and serotonin receptors, exhibiting a comparatively diminished affinity for histaminergic, cholinergic muscarinic, and α-adrenergic receptor subtypes (Ben-Levyet al., 1998). One of the exciting advances in drug discovery is the coordinated combination of several beneficial moieties on identical substances, especially for the treatment of certain illnesses (Chanet al., 2017). The ideal environment for creating novel compounds with pleiotropic pharmacological effects is in the area of pharmaceutical benefits that reduce the need for polypharmacy, such as treatments that also reduce iatrogenic side effects, prevent symptoms, or enhance beneficial effects through adjuvant therapeutic mechanisms (Cuadrado and Nebreda, 2010). Dual inhibitors frequently have high atomic weights, which may reduce the possibility of their therapeutic effects. As a result, pharmacokinetic properties and safety profiles need to be carefully considered while constructing dual inhibitors (Fieldet al., 2018). The combination of imidazole with benzene results in the heterocyclic aromatic organic molecule known as phenothiazine. Because of their many effects, they have been important in medical chemistry because of their analgesic, anti-tumour, antiviral, and psychotropic qualities (Gröberet al., 2013). The complex biochemical response of body tissues to infections, which is an immune cell defines mechanism (Hobbenaghiet al., 2013). Monoamine oxidase (PDB ID: 2BXR) Enzyme: (COX), which comes in the forms Monoamine oxidase (PDB ID: 2BXR) Enzyme: is essential protein needed for converting arachidonic acid to prostaglandins. Prostaglandins are supplied by COX-1 (PDB: 3KK6) while MONOAMINE OXIDASE (PDB ID: 2BXR) (PDB: 3LN1), is produced subsequent to inflammatory stimuli (Horiuchiet al., 1999). In human cancers, MONOAMINE OXIDASE (PDB ID: 2BXR) (PDB: 3LN1) contributes to cell division, angiogenesis, and death. Molecular docking, a method for determining the interaction between a protein and ligand, describes the orientation, binding interactions, and binding energy of the molecule (Kumaret al., 2003). Pharmacophore techniques are necessary to predict the common molecular traits of those involved within the molecular-level interactions involving biological targets (Lee & Young, 1996). The aim of this study was to analyse the physicochemical and ADMET properties of the synthesized drugs and identify the molecular interactions in COX and phenothiazine analogues (Ono & Han, 2000). Furthermore, unique characteristics were investigated through pharmacophore modelling studies. Given the claimed anti-depressant activity properties of phenothiazine derivatives, it is crucial to illustrate their mechanism of action (Rameshet al., 2014). To ascertain these compounds in silico inhibitory effect, were docked with Monoamine oxidase (PDB ID: 2BXR) Enzyme: In order to determine how phenothiazine function to inhibit Monoamine Oxidase (pdb id: 2bxr) enzymes, which is what gives them their Anti-Depressants Activity properties, in-silico research is necessary. An essential mechanism against all types of chemical, physical, and viral attacks is depression (Rathodet al., 2016). The body experiences pathological situations as organ rejection, autoimmune disorders, and allergies when this pathway is dysregulating (Streitet al., 2004). Since NSAID show lowering fever, Depression and pain, millions of patients use them all over the world. NSAIDs function pharmacologically by blocking the actions of Monoamine oxidase (PDB ID: 2BXR), which stops enzymatic biotransformation of arachidonic acid into related pro-inflammatory prostaglandins and Thromboxane (TXs). In 1997, Needleman and Isakson published the first description of two isoenzymes. The “House Keeping” enzyme, monoamine oxidase (PDB ID: 2BXR), Numerous things can cause the “inducible” enzyme to produce inflammatory reactions. The specific functions of the monoamine oxidase isoform (PDB ID: 2BXR), which is uniquely expressed in certain regions of the Central Nervous System (CNS) remain unclear. The second Monoamine oxidase (pdb id: 2bxr), produced in pathogenic and pro-inflammatory stimuli, including cytokines, lipopolysaccharides and phorbol esters. Monoamine oxidase (pdb id: 2bxr) enzyme, is responsible for both PGs release and the Depression. Monoamine oxidase (PDB ID: 2BXR) has been connected in numerous studies to a variety of clinical conditions, including depression, neurological diseases, and cancer. Hence, it is also used to treat cancer. Selective monoamine oxidase (PDB id: 2bxr) (PDB: 3LN1) inhibitor drugs, such as celecoxib, rofecoxib, and valdecoxib, were developed to lessen these severe adverse effects. These drugs have better gastrointestinal safety profiles and the same analgesic effects as non-selective monoamine oxidase (pdb id: 2bxr) inhibitors. Unfortunately, as they have adverse effects on Monoamine oxidase (pdb id: 2bxr) pathway, which includes an elevated risk of myocardial infarction and high blood pressure, valdecoxib and rofecoxib have both been removed from the market. The adverse side effects of valdecoxib and rofecoxib were linked to their different chemical compositions. Because of this, research is still being done to find selective antidepressant activity drugs that have better safety profiles than the NSAIDs currently on the market. Derivatives of inhibition were tested in the monoamine oxidase (PDB ID: 2BXR) enzyme. subsequently to elucidate their potential mode of action, the synthesized compounds docked into the monoamine oxidase (pdb id: 2bxr) enzyme site.
MATERIALS AND METHODS
Molecular docking and the synthesis of ligands
Test compounds PS1-PS16 structures and the phenothiazine scheme were provided in Figure 3 and Table 1 prior to molecular docking. Using the ArgusLab 4.0.1 software and the semi-empirical method, we optimized. Getting proteins ready We used a variety of different Monoamine oxidase (PDB ID: 2BXR) Enzyme, enzyme crystal structures via the RCSB for docking studies. The study examined the compounds’ ADMET, physicochemical properties, and drug similarity. PDB provided the monoamine oxidase ribbon composition (PDB ID: 2BXR). An enzyme 1.5.6 was the version of auto dock. The selection was Chain A. After removing the water, polar hydrogen with Gasteiger charges was added. The grid map was their choice. ChemDraw Ultra 12.0 was utilized to create the 2D formula for the compounds. Utilizing Avogadro software, energy consumption was reduced. Discovery Studio was used to visualize the reactions, and Auto dock Vina was utilized to implement the docking technique. Optimising the docking method involved comparing the crystal structures of Celecoxib (CEL) with the expected conformations of the docking statistics. In Figure 1, the Ribbon Structure Protein structure’s crystalline form was superimposed; and in Figure 2, a Ramachandran plot was exhibited.
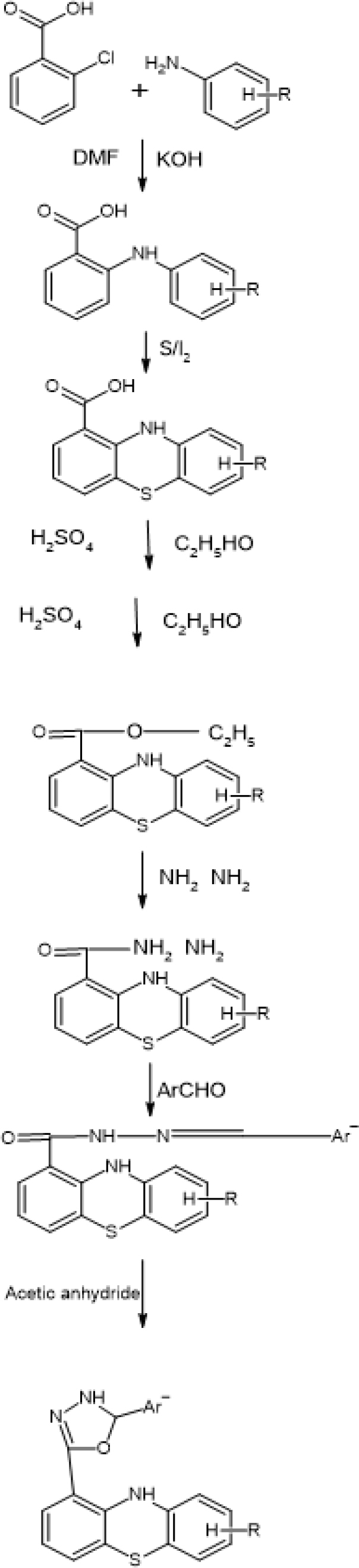
Figure 1:
Scheme of Phenothiazine.
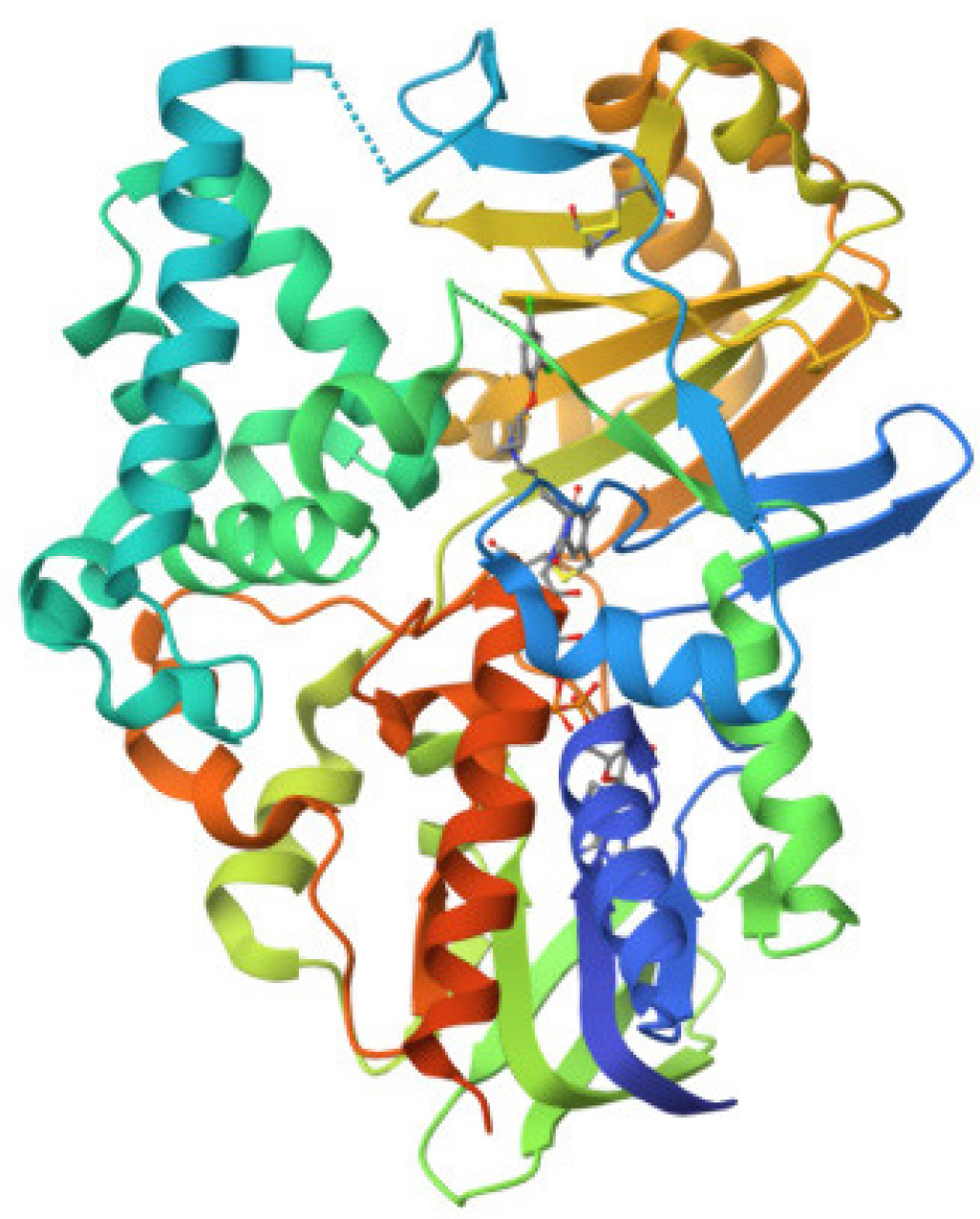
Figure 2:
3D Structure of monoamine oxidase (PDB ID: 2BXR).
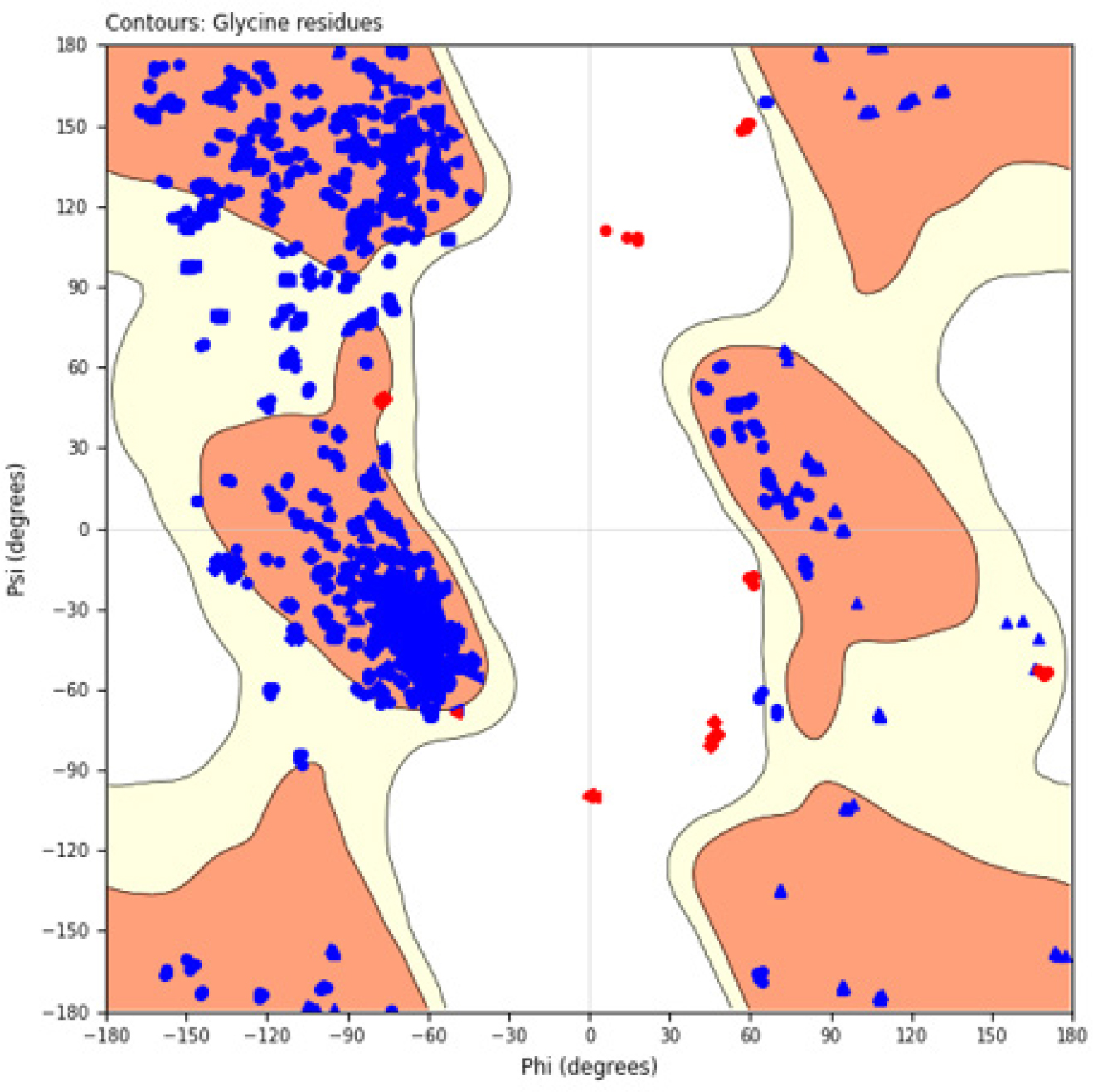
Figure 3:
Ramachandran Plot.
| Comp. Code | Ar | R | Comp.
Code |
Ar | R |
|---|---|---|---|---|---|
| PS1 | Cl | PS16 | Cl | ||
| PS2 | Cl | PS17 | Cl | ||
| PS3 | Cl | PS18 | Cl | ||
| PS4 | Cl | PS19 | Cl | ||
| PS5 | Cl | PS42 | Br | ||
| PS6 | Cl | PS44 | Br | ||
| PS7 | Cl | PS45 | Br | ||
| PS8 | Cl | PS46 | Br | ||
| PS9 | Cl | PS47 | NO2 | ||
| PS10 | Cl | PS51 | NO2 | ||
| PS11 | Cl | PS52 | NO2 | ||
| PS12 | Cl | PS53 | NO2 | ||
| PS13 | Cl | PS54 | NO2 | ||
| PS14 | Cl | PS62 | NO2 | ||
| PS15 | Cl | PS66 | NO2 |
RESULTS
Lipinski’s rule of five and molecular docking
Lipinski’s “rule of five”-a computational and experimental approach for calculating solubility and permeability-is followed throughout the drug development process. PDB ID for monoamine oxidase: 2BXR Enzyme: medicines and enzymes were docked, and the molecular interactions between the two were analysed. The 2D-3D Composition for molecular bindings is shown in Table 4 summarizes Chemical processes involving dynamic amino acid residues. While lead optimization and identification have historically been the main focus of Computer-Aided Molecular Design (CAMD), a number of innovative methods have been developed to help strengthen the binding affinities of drug candidates to particular receptors. These general guidelines evaluate a molecule’s drug-likeness and determine whether it possesses pharmacological action. The rule was found on the finding that the most of medications work when taken orally, a little lipophilic molecule. It is used when the pharmacologically active lead compound is iteratively enhanced to increase their activity and selectivity while preserving their drug-like physical and chemical properties in the process of creating novel medications. Rotation of bonds, Table 2 includes data on H-bond donors and acceptors, as well as violations of the Lipinski, Ghose, Veber, Egan, and Muegge rules. To make oral drugs that work, chemicals must have ADMET properties. A ligand’s toxicity is believed to be necessary for it to serve as an effective tool, and Qik-Prop generates descriptions that are physically significant. The ligand preparation tool used in research is the Ligprep module. The protein preparation wizard is used to prepare proteins. Monoamine oxidase’s X-ray crystal structures are obtaine from the PDB data bank an enzyme: Receptor Grid Generation Wizard-generated grid. Receptor Grid Generation Wizard were given in Figure 4 Glide XP coupled the ligand with the protein, and the interactions were seen. The scoring function allocates points based on the ideal ligand-protein interaction. The docking positions were evaluated in the extra-precision mode. The software recognizes hydrogen bonds, hydrophobic interactions, metal-ligation interactions, and steric conflicts. Every material has a molecular weight that is less than 500, ranging from 400 to 500. The calculated log p values of the compounds range from 3.56 to 5.35. The materials under investigation contain hydrogen bond donors.
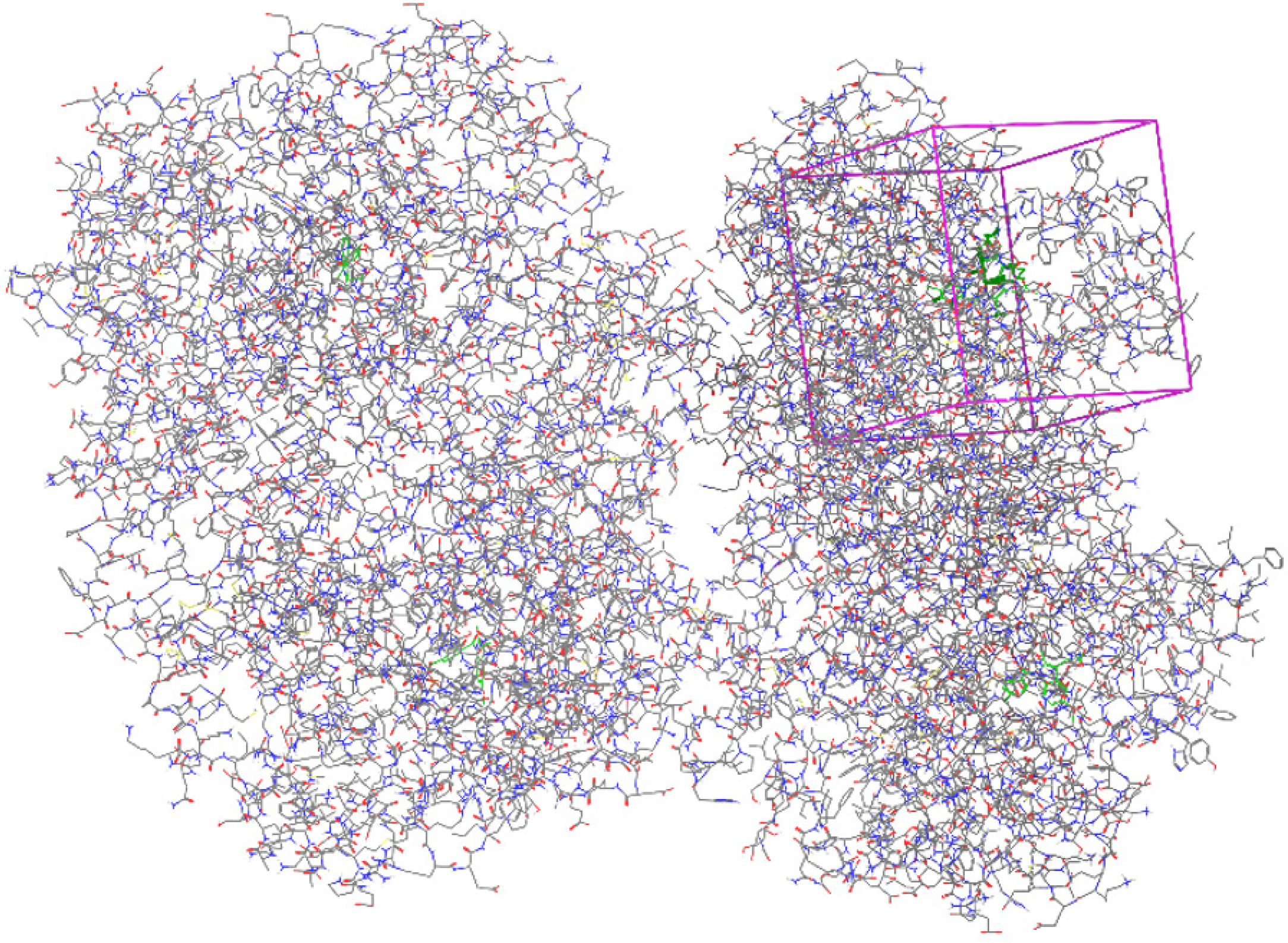
Figure 4:
Receptor Grid Generation Wizard.
| Compound.
Code |
Formula | Lipinski rule of five | Viber’s rule | |||||
|---|---|---|---|---|---|---|---|---|
| MW | iLOGP | #H-bond acceptors | #H-bond donors | Lipinski #violations | Rotatable bonds | TPSA | ||
| PS 1 | C21H16ClN3OS | 393.89 | 3.65 | 2 | 2 | 1 | 2 | 70.95 |
| PS 2 | C22H18ClN3OS | 407.92 | 3.7 | 2 | 2 | 1 | 3 | 70.95 |
| PS 3 | C20H14ClN3O2S | 395.86 | 2.98 | 3 | 3 | 1 | 2 | 91.18 |
| PS 4 | C23H20ClN3OS | 421.94 | 3.87 | 2 | 2 | 1 | 4 | 70.95 |
| PS 5 | C24H22ClN3OS | 435.97 | 3.88 | 2 | 2 | 1 | 5 | 70.95 |
| PS 6 | C21H16ClN3O2S | 409.89 | 3.49 | 3 | 3 | 1 | 3 | 91.18 |
| PS 7 | C23H20ClN3O2S | 437.94 | 3.34 | 3 | 3 | 1 | 5 | 91.18 |
| PS 8 | C22H18ClN3O2S | 423.92 | 3.39 | 3 | 3 | 1 | 4 | 91.18 |
| PS 9 | C21H14ClN3O3S | 423.87 | 2.65 | 4 | 3 | 1 | 3 | 108.25 |
| PS 10 | C21H14ClN3O3S | 423.87 | 2.89 | 4 | 3 | 1 | 3 | 108.25 |
| PS 11 | C21H14ClN3O3S | 423.87 | 2.8 | 4 | 3 | 1 | 3 | 108.25 |
| PS 12 | C20H13ClN4O3S | 424.86 | 2.85 | 4 | 2 | 1 | 3 | 116.77 |
| PS 13 | C20H13ClN4O3S | 424.86 | 2.7 | 4 | 2 | 1 | 3 | 116.77 |
| PS 14 | C20H13ClN4O3S | 424.86 | 2.87 | 4 | 2 | 1 | 3 | 116.77 |
| PS 15 | C20H13Cl2N3OS | 414.31 | 3.45 | 2 | 2 | 1 | 2 | 70.95 |
| PS 16 | C20H13Cl2N3OS | 414.31 | 3.63 | 2 | 2 | 1 | 2 | 70.95 |
| PS 17 | C20H13Cl2N3OS | 414.31 | 3.7 | 2 | 2 | 1 | 2 | 70.95 |
| PS 18 | C26H18ClN3OS | 455.96 | 3.73 | 2 | 2 | 1 | 3 | 70.95 |
| PS 19 | C20H14ClN3OS | 379.86 | 3.6 | 2 | 2 | 1 | 2 | 70.95 |
| PS 20 | C26H18ClN3OS | 455.96 | 3.77 | 2 | 2 | 1 | 3 | 70.95 |
| PS 21 | C22H16ClN3O2S | 421.9 | 3.16 | 3 | 2 | 1 | 3 | 88.02 |
| PS 22 | C22H16ClN3O2S | 421.9 | 3.32 | 3 | 2 | 1 | 3 | 88.02 |
| PS 23 | C22H16ClN3O2S | 421.9 | 3.3 | 3 | 2 | 1 | 3 | 88.02 |
| PS 24 | C21H16BrN3OS | 438.34 | 3.85 | 2 | 2 | 1 | 2 | 70.95 |
| PS 25 | C22H18BrN3OS | 452.37 | 3.78 | 2 | 2 | 1 | 3 | 70.95 |
| PS 26 | C20H14BrN3O2S | 440.31 | 3 | 3 | 3 | 1 | 2 | 91.18 |
| PS 27 | C23H20BrN3OS | 466.39 | 4.03 | 2 | 2 | 1 | 4 | 70.95 |
| PS 28 | C24H22BrN3OS | 480.42 | 4.27 | 2 | 2 | 1 | 5 | 70.95 |
| PS 29 | C21H16BrN3O2S | 409.89 | 3.49 | 3 | 3 | 1 | 3 | 91.18 |
| PS 30 | C23H20BrN3O2S | 482.39 | 3.55 | 3 | 3 | 1 | 5 | 91.18 |
| PS 31 | C22H18BrN3O2S | 468.37 | 3.5 | 3 | 3 | 1 | 4 | 91.18 |
| PS 32 | C21H14BrN3O3S | 468.32 | 2.93 | 4 | 3 | 1 | 3 | 108.25 |
| PS 33 | C21H14BrN3O3S | 468.32 | 2.69 | 4 | 3 | 1 | 3 | 108.25 |
| PS 34 | C21H14BrN3O3S | 468.32 | 2.69 | 4 | 3 | 1 | 3 | 108.25 |
| PS 35 | C20H13BrN4O3S | 469.31 | 2.98 | 4 | 2 | 1 | 3 | 116.77 |
| PS 36 | C20H13BrN4O3S | 469.31 | 2.8 | 4 | 2 | 1 | 3 | 116.77 |
| PS 37 | C20H13BrN4O3S | 469.31 | 2.94 | 4 | 2 | 1 | 3 | 116.77 |
| PS 38 | C20H13BrClN3OS | 458.76 | 3.58 | 2 | 2 | 1 | 2 | 70.95 |
| PS 39 | C20H13BrClN3OS | 458.76 | 3.68 | 2 | 2 | 1 | 2 | 70.95 |
| PS 40 | C20H14Br2ClN3OS | 539.67 | 0 | 2 | 2 | 2 | 2 | 70.95 |
| PS 41 | C26H19Br2N3OS | 581.32 | 0 | 2 | 2 | 2 | 3 | 70.95 |
| PS 42 | C20H14BrN3OS | 424.31 | 3.71 | 2 | 2 | 1 | 2 | 70.95 |
| PS 43 | C26H18BrN3OS | 500.41 | 3.8 | 2 | 2 | 2 | 3 | 70.95 |
| PS 44 | C22H16BrN3O2S | 466.35 | 3.27 | 3 | 2 | 1 | 3 | 88.02 |
| PS 45 | C22H16BrN3O2S | 466.35 | 3.38 | 3 | 2 | 1 | 3 | 88.02 |
| PS 46 | C22H16BrN3O2S | 466.35 | 3.39 | 3 | 2 | 1 | 3 | 88.02 |
| PS 47 | C22H17N3O3S | 403.45 | 2.99 | 3 | 2 | 0 | 3 | 104.41 |
| PS 48 | C23H19N3O3S | 417.48 | 3.24 | 3 | 2 | 1 | 4 | 104.41 |
| PS 49 | C24H21N3O3S | 431.51 | 3.43 | 3 | 2 | 1 | 5 | 104.41 |
| PS 50 | C25H23N3O3S | 445.53 | 3.83 | 3 | 2 | 1 | 6 | 104.41 |
| PS 51 | C24H21N3O4S | 447.51 | 3.18 | 4 | 3 | 0 | 6 | 124.64 |
| PS 52 | C23H19N3O4S | 433.48 | 3.02 | 4 | 3 | 0 | 5 | 124.64 |
| PS 53 | C22H17N3O4S | 419.45 | 2.87 | 4 | 3 | 0 | 4 | 124.64 |
| PS 54 | C21H15N3O4S | 405.43 | 2.16 | 4 | 3 | 0 | 3 | 124.64 |
| PS 55 | C22H15N3O5S | 433.44 | 2.55 | 5 | 3 | 0 | 4 | 141.71 |
| PS 56 | C22H15N3O5S | 433.44 | 2.26 | 5 | 3 | 0 | 4 | 141.71 |
| PS 57 | C22H15N3O5S | 433.44 | 1.93 | 5 | 3 | 0 | 4 | 141.71 |
| PS 58 | C21H14N4O5S | 434.42 | 2.55 | 5 | 2 | 0 | 4 | 150.23 |
| PS 59 | C21H14N4O5S | 434.42 | 2.65 | 5 | 2 | 0 | 4 | 150.23 |
| PS 60 | C21H14N4O5S | 434.42 | 2.58 | 5 | 2 | 0 | 4 | 150.23 |
| PS 61 | C21H14ClN3O3S | 423.87 | 2.98 | 3 | 2 | 1 | 3 | 104.41 |
| PS 62 | C21H14ClN3O3S | 423.87 | 2.9 | 3 | 2 | 1 | 3 | 104.41 |
| PS63 | C23H17N3O4S | 431.46 | 2.9 | 4 | 2 | 0 | 4 | 121.48 |
| PS 64 | C23H17N3O4S | 431.46 | 2.7 | 4 | 2 | 0 | 4 | 121.48 |
| PS 65 | C23H17N3O4S | 431.46 | 2.66 | 4 | 2 | 0 | 4 | 121.48 |
| PS 66 | C21H14ClN3O3S | 423.87 | 2.77 | 3 | 2 | 1 | 3 | 104.41 |
| PS 67 | C27H19N3O3S | 465.52 | 3.73 | 3 | 2 | 1 | 4 | 104.41 |
| PS 68 | C27H19N3O3S | 465.52 | 3.57 | 3 | 2 | 1 | 4 | 104.41 |
| PS 69 | C21H15N3O3S | 389.43 | 2.9 | 3 | 2 | 0 | 3 | 104.41 |
Silico Admet
Quantum approaches are used in molecular modelling to assess the likelihood of interactions, such as those involving cytochrome P450, are involved in ADME processes. Data modeling frequently uses QSAR techniques. These employ statistical techniques to find correlations in a group of structural molecules, such as GI absorption, BBB penetration, P-glycoprotein protein, CYP1A2 inhibitor, CYP2C9 inhibitor, CYP2D6 inhibitor, and CYP3A4 inhibitor, and specific properties. Models for effective prediction for the ADMET parameters require the selection of a suitable mathematical approach, the use of suitable chemical descriptors for the ADMET endpoint, and a sufficiently large collection of data related to the same endpoint for model validation. Effective property prediction methods are becoming more important to accomplish two primary objectives. First, novel compound libraries should be created to reduce the risk. Second, the most promising compounds should be the focus of screening and testing. Predicting the goal of this study is to find features that provide information regarding dosage quantity and frequency, such as oral absorption, bioavailability, BBB penetration, clearance, and Vd (for frequency). Two sorts of computational approaches are used: data modelling and molecular modelling. This article examines recent advances in the prediction of physicochemical properties relevant to ADME features (like absorption), and toxicity issues see Table 3. The use of automated medium and HTS in vitro tests will increase throughout the next 10 or more years.
| Comp
code |
GI absorption | BBB permeant | Pgp substrate | CYP1A2 inhibitor | CYP2C19 inhibitor | CYP2C9 inhibitor | CYP2D6 inhibitor | CYP3A4 inhibitor | Lipinski #violations | Ghose | Veber | Egan | Muegge | Bioavailability Score |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| PS 1 | High | Yes | Yes | Yes | Yes | Yes | No | Yes | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 2 | High | Yes | Yes | Yes | Yes | Yes | No | Yes | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 3 | High | No | No | No | Yes | Yes | No | Yes | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 4 | High | No | Yes | Yes | Yes | Yes | Yes | Yes | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 5 | High | No | Yes | Yes | Yes | Yes | Yes | Yes | 1 | 1 | 0 | 0 | 1 | 0.55 |
| PS 6 | High | No | Yes | Yes | Yes | Yes | Yes | Yes | 1 | 0 | 0 | 0 | 0 | 0.55 |
| PS 7 | High | No | Yes | Yes | Yes | Yes | Yes | Yes | 1 | 1 | 0 | 0 | 1 | 0.55 |
| PS 8 | High | No | Yes | Yes | Yes | Yes | Yes | Yes | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 9 | High | No | No | No | No | Yes | No | No | 1 | 0 | 0 | 0 | 1 | 0.56 |
| PS 10 | High | No | No | No | No | Yes | No | No | 1 | 0 | 0 | 0 | 1 | 0.56 |
| PS 11 | High | No | No | No | No | Yes | No | No | 1 | 0 | 0 | 0 | 1 | 0.56 |
| PS 12 | High | No | No | No | Yes | Yes | No | No | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 13 | High | No | No | No | Yes | Yes | No | No | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 14 | High | No | No | No | Yes | Yes | No | No | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 15 | High | Yes | Yes | Yes | Yes | Yes | No | No | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 16 | High | Yes | Yes | Yes | Yes | Yes | No | No | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 17 | High | Yes | No | Yes | Yes | Yes | No | No | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 18 | High | No | No | Yes | Yes | Yes | No | No | 1 | 2 | 0 | 0 | 1 | 0.55 |
| PS 19 | High | Yes | No | Yes | Yes | Yes | No | Yes | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 20 | High | No | No | Yes | Yes | Yes | No | No | 1 | 2 | 0 | 0 | 1 | 0.55 |
| PS 21 | High | No | No | No | Yes | Yes | No | Yes | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 22 | High | No | No | Yes | Yes | Yes | No | Yes | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 23 | High | No | No | Yes | Yes | Yes | No | Yes | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 24 | High | Yes | Yes | Yes | Yes | Yes | No | No | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 25 | High | Yes | Yes | Yes | Yes | Yes | No | Yes | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 26 | High | No | No | No | No | Yes | No | Yes | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 27 | High | No | Yes | Yes | Yes | Yes | Yes | Yes | 1 | 1 | 0 | 0 | 1 | 0.55 |
| PS 28 | High | No | Yes | Yes | No | Yes | Yes | Yes | 1 | 2 | 0 | 0 | 1 | 0.55 |
| PS 29 | High | No | Yes | Yes | Yes | Yes | Yes | Yes | 1 | 0 | 0 | 0 | 0 | 0.55 |
| PS 30 | High | No | Yes | Yes | No | Yes | Yes | Yes | 1 | 2 | 0 | 0 | 1 | 0.55 |
| PS 31 | High | No | Yes | Yes | No | Yes | Yes | Yes | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 32 | High | No | No | No | No | Yes | No | No | 1 | 0 | 0 | 0 | 1 | 0.56 |
| PS 33 | High | No | No | No | No | Yes | No | No | 1 | 0 | 0 | 0 | 1 | 0.56 |
| PS 34 | High | No | No | No | No | Yes | No | No | 1 | 0 | 0 | 0 | 1 | 0.56 |
| PS 35 | High | No | No | No | Yes | Yes | No | No | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 36 | High | No | No | No | Yes | Yes | No | No | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 37 | High | No | No | No | Yes | Yes | No | No | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 38 | High | No | No | Yes | Yes | Yes | No | No | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 39 | High | No | No | Yes | Yes | Yes | No | No | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 40 | High | No | Yes | Yes | No | No | No | No | 2 | 3 | 0 | 0 | 1 | 0.17 |
| PS 41 | Low | No | Yes | Yes | Yes | No | No | No | 2 | 3 | 0 | 1 | 1 | 0.17 |
| PS 42 | High | Yes | No | Yes | Yes | Yes | No | Yes | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 43 | High | No | No | Yes | Yes | No | No | No | 2 | 3 | 0 | 0 | 1 | 0.17 |
| PS 44 | High | No | No | Yes | Yes | Yes | No | Yes | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 45 | High | No | No | Yes | Yes | Yes | No | Yes | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 46 | High | No | No | Yes | Yes | Yes | No | Yes | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 47 | High | No | Yes | Yes | Yes | Yes | No | Yes | 0 | 0 | 0 | 0 | 1 | 0.55 |
| PS 48 | High | No | No | Yes | Yes | Yes | No | Yes | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 49 | High | No | No | Yes | Yes | Yes | No | Yes | 1 | 1 | 0 | 0 | 1 | 0.55 |
| PS 50 | Low | No | Yes | Yes | Yes | Yes | Yes | Yes | 1 | 2 | 0 | 0 | 1 | 0.55 |
| PS 51 | Low | No | Yes | Yes | Yes | Yes | Yes | Yes | 0 | 1 | 0 | 0 | 0 | 0.55 |
| PS 52 | High | No | Yes | Yes | Yes | Yes | Yes | Yes | 0 | 0 | 0 | 0 | 0 | 0.55 |
| PS 53 | High | No | Yes | Yes | Yes | Yes | No | Yes | 0 | 0 | 0 | 0 | 0 | 0.55 |
| PS 54 | High | No | Yes | Yes | Yes | Yes | No | Yes | 0 | 0 | 0 | 0 | 0 | 0.55 |
| PS 55 | Low | No | No | Yes | Yes | Yes | No | No | 0 | 0 | 1 | 1 | 0 | 0.56 |
| PS 56 | Low | No | No | Yes | Yes | Yes | No | No | 0 | 0 | 1 | 1 | 0 | 0.56 |
| PS 57 | Low | No | No | Yes | Yes | Yes | No | No | 0 | 0 | 1 | 1 | 0 | 0.56 |
| PS 58 | Low | No | No | Yes | Yes | Yes | No | Yes | 0 | 0 | 1 | 1 | 1 | 0.55 |
| PS 59 | Low | No | No | No | Yes | Yes | No | No | 0 | 0 | 1 | 1 | 1 | 0.55 |
| PS 60 | Low | No | No | Yes | Yes | Yes | No | Yes | 0 | 0 | 1 | 1 | 1 | 0.55 |
| PS 61 | High | No | No | Yes | Yes | Yes | No | Yes | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 62 | High | No | No | Yes | Yes | Yes | No | Yes | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS63 | Low | No | No | Yes | Yes | Yes | No | Yes | 0 | 0 | 0 | 0 | 0 | 0.55 |
| PS 64 | Low | No | No | Yes | Yes | Yes | No | Yes | 0 | 0 | 0 | 0 | 0 | 0.55 |
| PS 65 | Low | No | No | Yes | Yes | Yes | No | Yes | 0 | 0 | 0 | 0 | 0 | 0.55 |
| PS 66 | High | No | No | Yes | Yes | Yes | No | Yes | 1 | 0 | 0 | 0 | 1 | 0.55 |
| PS 67 | Low | No | No | No | Yes | Yes | No | No | 1 | 2 | 0 | 1 | 1 | 0.55 |
| PS 68 | Low | No | No | No | Yes | Yes | No | No | 1 | 2 | 0 | 1 | 1 | 0.55 |
| PS 69 | High | No | Yes | Yes | Yes | Yes | No | Yes | 0 | 0 | 0 | 0 | 1 | 0.55 |
DISCUSSION
The optimal conformation gets determined by analogs is the good interactions and Auto dock values that are close to zero. Both complexes’ Ramachandran Plot RMSD plot analysis revealed that the PS9-protein complex had a stable trajectory for more research after achieving excellent stability at 100 ns. Analysis of the synthetic phenothiazine derivatives. According to data all “compounds had molecular weights less than 500,” show that they have an action site binding effect. A favorable in vivo reaction requires a balance between pharmacodynamics and pharmacokinetic parameters. Additionally, ADMET provides additional information on medication dosage and regimen. In accordance with ” An oral medication is selected according to Lipinski’s “rule of five” if its log p value is less than five, its molecular weight is less than 500, and its hydrogen bond donors and acceptors are fewer than five and ten, respectively. Rotatable bond count indicates molecular flexibility, which is essential for oral bioavailability. Given its indirect relationship to percentage absorption, it was also proposed that TPSA be used as a 3D descriptor of the quantity of hydrogen bonding groups. Each drug had a Log p value less than 5, showing better absorption and penetration of cell membranes. The “binding energies and hydrogen bonds” of PS1-PS16 are detailed for several phenothiazine derivatives in Table 1. Furthermore, TPSA was suggested as a 3D descriptor for the number of hydrogen bonding groups due to its indirect relationship to percentage absorption. A Log p value less than five is signified that each medication had superior cell membrane penetration and absorption. Table 1 lists the various phenothiazine derivatives along with information on the “binding energies and hydrogen bonds” of PS1-PS16. Phe381, Leu-384, Tyr-385, Trp-387, Phe-518, Gly-526, Met-522, Tyr348, Val-349, Leu-352 Monoamine included active amino acids. oxidase (PDB ID: 2BXR) Enzyme: PS5 and PS6 demonstrated Pi-Pi stacking to TYR355: HH; SER 530: HG; SER530: HB1; TYR355; PHE381; TYR385; TRP387; TRP387; TRP387; ALA527: N; LEU384; MET522; VAL349; ALA527; and LEU531 across the phenothiazine and form an H-bond with Ser-530 through the phenothiazine ring’s nitrogen. PS15 showed H- bonding with Ser530 and Pi-Pi stacking with Tyr385 across the phenothiazine ring. PS11 demonstrated Pi-Pi stacking with ARG120:HH11; ARG120:HH12; TYR 355: HH; ARG120:NH1; ARG120:NH2; VAL89; VAL116; VAL349; ALA527; VAL349 and LEU352; ALA527 via benzene ring and H-bond with Met-522 via nitrogen. A reasonable docking score of -8.572 kcal/mol was obtained for PS15 and PS11 when comparing the 16 phenothiazine derivatives to the standard Indomethacin (-10.705 kcal/mol). ARG120:HH11; ARG120:HH12; TYR355: HH; ARG120:NH1; ARG120:NH2; VAL 89; VAL116; VAL349; ALA527; VAL349 and LEU352; ALA527, the monoamine oxidase enzyme contained active amino acids. (PDB ID: 2BXR) Enzyme: enzymes. PS5 demonstrated H-Bond with TYR355: HH; SER 530: HG; SER530:HB1; TYR355; PHE381; TYR385; TRP387; TRP387; TRP387; ALA527: N; LEU384; MET522; VAL349; ALA527; and LEU531 nitrogen of phenothiazine and Pi-Pi stacking with Tyr-385 via benzene. PS13 demonstrated Pi-cation interaction to TYR 355:HH; SER530: HG; SER530:HB1; PHE381; TYR385; TRP387; TRP387; TRP387; VAL116; LEU359; LEU531; LEU384; MET522; VAL349; ALA;527 and LEU531 via phenothiazine and Pi-pi stacking with Tyr-385 and Trp-387 via benzene. PS9 demonstrated H-bond with ARG120:HH12; ARG120:NH1; LEU93; VAL116; VAL116; VAL349; ALA527; LEU352 and Ser-530 via phenothiazine and Pi-Pi stacking with Tyr-385. When compared to the conventional Indomethacin (-10.099 kcal/mol), 16 phenothiazine derivatives, PS5, PS6, PS11, PS13, and PS15 displayed higher docking ranging from -8.25 to -8.51 kcal/mol (Table 5). The binding affinity score of compound PS5 with COX-1 (PDB: 3KK6) is -56.79 kcal/mol, while compound PS6 has a binding affinity score of -60.27 kcal/mol with MONOAMINE OXIDASE (PDB ID: 2BXR) (PDB: 3LN1). High ADME was also indicated by the compounds’ oral absorption percentage, which varied from 70.69 to 73.87%. Rotatable bonds ranged from 7 to 8 (<10), and their respective TPSA values were 101.3 and 104.80 A2 (140 A2). It is often believed that a molecule has both lipophilicity and hydrophilicity if it is soluble in water and meets Lipinski’s requirements. The enzymes’ interactions with the monoamine oxidase (PDB ID: 2BXR) enzyme demonstrated their anti-depressant activity. TYR355: HH; SER530:HG; SER530:HB1; TYR355; PHE381; TYR385; TRP387; TRP387; TRP387; ALA527:N; LEU384; MET522; VAL349; ALA527; and LEU531 were monoamine oxidase’s active amino acids (PDB ID: 2BXR). Enzyme: PS13 and PS15 showed that they could form an H-bond with Ser-530 through the nitrogen of the phenothiazine ring and Pi-Pi stack to Tyr385 and Trp387 via the phenothiazin.PS12 demonstrated Pi-Pi stacking with TYR355: HH; SER530:HG; SER530:HB1; TYR355; PHE381; TYR385; TRP387; TRP387; TRP387; ALA527:N; VAL349; ALA527; and LEU531 via the phenothiazin ring and hydrogen bonding with Ser530. PS14 demonstrated Pi-Pi stacking with Tyr-385 via benzene ring and H-bond with Met-522 via nitrogen. Comparing the 16 phenothiazin derivatives to the standard Indomethacin (- 10.705 kcal/mol), A reasonable docking score of -8.572 kcal/mol was obtained by PS14 and PS15.Phe381, Leu384, Tyr385, Trp387, were amino acids that were active in the monoamine oxidase enzyme (PDB ID: 2BXR). Enzyme: the enzymes. Pi-Pi stacking with Tyr-385 via benzene and H-Bond with Ser-530 nitrogen of phenothiazine were proven by BI5. PS10 demonstrated Pi-cation interaction to TYR355: HH; SER530:HG; SER530:HB1; TYR355; PHE381; TYR385; TRP387; TRP387; TRP387; ALA527:N; LEU384; MET522; VAL349; ALA527; and LEU531 via phenothiazine and Pi-pi stacking with Tyr-385 and Trp-387 via benzene. PS1 demonstrated H-bond with TYR355: HH; SER530:HG; SER530:HB1; TYR355; PHE381; TYR385; TRP387; TRP387; TRP387; ALA527:N; LEU384; MET522; VAL349; ALA527; and LEU531 via phenothiazine and Pi-Pi stacking with Tyr-385. A balanced combination of pharmacodynamics and pharmacokinetic properties is necessary for a favourable reaction in vivo. ADMET also provides more information on medication dosage and regimen.
| Compound
Code |
Active Amino Acid | Bond Type | Docking Score | 2D | 3D |
|---|---|---|---|---|---|
| PS1 | ASP141
GLU458 TRP144 LEU460 TYR35 ARG454 ARG457 |
Conventional Hydrogen Bond
Pi-Anion Pi-Pi T-shaped Alkyl Pi-Alkyl Pi-Alkyl Pi-Alkyl |
-10.1 | ||
| PS2 | ASP141
ARG457 GLU458 ASN461 TRP144 LEU460 TYR35 ARG454 ARG457 |
Conventional Hydrogen Bond
Carbon Hydrogen Bond Pi-Anion Pi-Donor Hydrogen Bond Pi-Pi T-shaped Alkyl Pi-Alkyl Pi-Alkyl Pi-Alkyl |
-10.1 | ||
| PS3 | GLN425
GLU458 ASN461 ARG457 |
Conventional Hydrogen Bond
Pi-Anion Pi-Donor Hydrogen Bond Pi-Alkyl |
-9.9 | ||
| PS4 | GLU216
GLU216 CYS406 TYR407 TRP441 TYR69 GLY66 PRO72 VAL303 LYS305 CYS406 PHE352 TRP397 TRP441 |
Pi-Anion
Pi-Anion Pi-Sulfur Pi-Pi Stacked Pi-Pi Stacked Pi-Pi T-shaped Amide-Pi Stacked Alkyl Alkyl Alkyl Alkyl Pi-Alkyl Pi-Alkyl Pi-Alkyl |
-11.3 | ||
| PS5 | SER209
GLU216 PHE208 PHE208 TYR407 TYR444 ARG51 LEU97 CYS323 LEU337 ARG51 MET445 ILE335 LEU337 |
Conventional Hydrogen Bond
Pi-Anion Pi-Pi Stacked Pi-Pi Stacked Pi-Pi Stacked Pi-Pi T-shaped Alkyl Alkyl Alkyl Alkyl Alkyl Alkyl Pi-Alkyl Pi-Alkyl |
-10 | ||
| PS6 | ASP141
GLN425 ARG457 GLU458 ASN461 TRP144 ARG457 |
Conventional Hydrogen Bond
Conventional Hydrogen Bond Carbon Hydrogen Bond Pi-Anion Pi-Donor Hydrogen Bond Pi-Pi T-shaped Pi-Alkyl |
-10.1 | ||
| PS7 | ARG457
ARG424 GLU458 ASN461 GLU458 ARG457 |
Conventional Hydrogen Bond
Conventional Hydrogen Bond Conventional Hydrogen Bond Pi-Donor Hydrogen Bond Pi-Sigma Pi-Alkyl |
-9.8 | ||
| PS8 | LEU277
TYR402 LYS280 ALA44 ILE273 TYR402 HIS282 LYS280 LYS280 LEU277 |
Conventional Hydrogen Bond
Carbon Hydrogen Bond Pi-Cation Pi-Sigma Pi-Sigma Pi-Pi T-shaped Pi-Alkyl Pi-Alkyl Pi-Alkyl Pi-Alkyl |
-10.5 | ||
| PS9 | THR439
THR439 LYS440 LYS440 LYS440 LYS199 LYS440 LYS440 LYS440 |
Conventional Hydrogen Bond
Sulfur-X Pi-Cation Pi-Sigma Pi-Sigma Alkyl Pi-Alkyl Pi-Alkyl Pi-Alkyl |
-8.7 | ||
| PS10 | ARG424
ASP141 GLU458 ARG454 ARG457 ARG457 LEU460 |
Conventional Hydrogen Bond
Conventional Hydrogen Bond Pi-Anion Pi-Alkyl Pi-Alkyl Pi-Alkyl Pi-Alkyl |
-10.3 | ||
| PS11 | ARG424
PRO426 GLU458 GLU458 GLU458 ARG424 PRO426 ARG424 |
Conventional Hydrogen Bond
Carbon Hydrogen Bond Pi-Anion Pi-Anion Pi-Anion Alkyl Pi-Alkyl Pi-Alkyl |
-10.1 | ||
| PS12 | ALA68
SER209 GLU216 ILE335 LEU337 PHE208 PHE208 TYR407 TYR444 LEU97 CYS323 LEU337 |
Conventional Hydrogen Bond
Sulfur-X Pi-Anion Pi-Sigma Pi-Sigma Pi-Pi Stacked Pi-Pi Stacked Pi-Pi Stacked Pi-Pi T-shaped Alkyl Alkyl Alkyl |
-11.3 | ||
| PS13 | ASP141
GLU458 ASN461 TRP144 ARG457 |
Conventional Hydrogen Bond
Pi-Anion Pi-Donor Hydrogen Bond Pi-Pi T-shaped Pi-Alkyl |
-10.4 | ||
| PS14 | TRP144
TYR35 ASP141 ARG457 GLU458 ASN461 ARG454 |
Conventional Hydrogen Bond
Conventional Hydrogen Bond Conventional Hydrogen Bond Carbon Hydrogen Bond Pi-Anion Pi-Donor Hydrogen Bond Pi-Alkyl |
-10.5 | ||
| PS15 | ASP141
GLU458 ASN461 ARG457 |
Conventional Hydrogen Bond
Pi-Anion Pi-Donor Hydrogen Bond Pi-Alkyl |
-10.0 | ||
| PS16 | ASP141
GLU458 ASN461 LEU460 ARG457 |
Conventional Hydrogen Bond
Pi-Anion Pi-Donor Hydrogen Bond Alkyl Pi-Alkyl |
-10.0 | ||
| PS17 | ASN461
ARG421 GLU458 ARG424 LEU460 ARG421 LEU460 |
Conventional Hydrogen Bond
Conventional Hydrogen Bond Pi-Anion Pi-Donor Hydrogen Bond Alkyl Pi-Alkyl Pi-Alkyl |
-10.3 | ||
| PS18 | SER209
GLU216 CYS406 PHE208 3PHE208 TYR407 TYR444 LEU97 CYS323 LEU337 ILE335 LEU337 ARG51 |
Conventional Hydrogen Bond
Pi-Anion Pi-Sulfur Pi-Pi Stacked Pi-Pi Stacked Pi-Pi Stacked Pi-Pi T-shaped Alkyl Alkyl Alkyl Pi-Alkyl Pi-Alkyl Pi-Alkyl |
-12.3 | ||
| PS19 | GLU216
ILE335 LEU337 PHE208 PHE208 TYR407 TYR444 CYS323 ILE335 LEU337 |
Pi-Anion
Pi-Sigma Pi-Sigma Pi-Pi Stacked Pi-Pi Stacked Pi-Pi Stacked Pi-Pi T-shaped Alkyl Alkyl Alkyl |
-11.1 | ||
| PS42 | SER209
GLU216 ILE335 PHE208 PHE208 TYR407 TYR444 LEU97 CYS323 LEU337 LEU337 |
Conventional Hydrogen Bond
Pi-Anion Pi-Sigma Pi-Pi Stacked Pi-Pi Stacked Pi-Pi Stacked Pi-Pi T-shaped Alkyl Alkyl Alkyl Pi-Alkyl |
-11.1 | ||
| PS44 | ARG424
ARG424 GLU458 ARG424 GLU458 PRO426 ARG424 |
Conventional Hydrogen Bond
Conventional Hydrogen Bond Conventional Hydrogen Bond Pi-Donor Hydrogen Bond Pi-Sigma Pi-Alkyl Pi-Alkyl |
-10.2 | ||
| PS45 | ASP141
ARG457 GLU458 ASN461 TRP144 ARG457 |
Conventional Hydrogen Bond
Carbon Hydrogen Bond Pi-Anion Pi-Donor Hydrogen Bond Pi-Pi T-shaped Pi-Alkyl |
-10.6 | ||
| PS46 | GLU216
ILE335 LEU337 PHE208 PHE208 TYR407 TYR444 CYS323 ILE335 LEU337 |
Pi-Anion
Pi-Sigma Pi-Sigma Pi-Pi Stacked Pi-Pi Stacked Pi-Pi Stacked Pi-Pi T-shaped Alkyl Alkyl Alkyl |
-12.0 | ||
| PS47 | ARG76
ARG421 ASP141 LEU460 TYR35 ARG424 ARG457 ARG454 ARG457 LEU460 |
Conventional Hydrogen Bond
Conventional Hydrogen Bond Pi-Anion Alkyl Pi-Alkyl Pi-Alkyl Pi-Alkyl Pi-Alkyl Pi-Alkyl Pi-Alkyl |
-10.1 | ||
| PS51 | ASN461
ARG424 GLU145 ARG457 GLU458 ASN461 ARG457 ARG457 ARG424 ARG457 |
Conventional Hydrogen Bond
Conventional Hydrogen Bond Conventional Hydrogen Bond Carbon Hydrogen Bond Pi-Anion Pi-Donor Hydrogen Bond Amide-Pi Stacked Pi-Alkyl Pi-Alkyl Pi-Alkyl |
-9.8 | ||
| PS52 | TYR35
ASN461 GLU145 GLU458 GLU458 ASN461 ARG454 ARG454 ARG457 |
Conventional Hydrogen Bond
Conventional Hydrogen Bond Conventional Hydrogen Bond Carbon Hydrogen Bond Pi-Anion Pi-Donor Hydrogen Bond Pi-Alkyl Pi-Alkyl Pi-Alkyl |
-9.6 | ||
| PS53 | SER209
GLY71 GLU216 GLU216 CYS406 TYR407 TRP441 GLY66,GLY67 |
Conventional Hydrogen Bond
Carbon Hydrogen Bond Pi-Anion Pi-Anion Pi-Sulfur Pi-Pi Stacked Pi-Pi Stacked Amide-Pi Stacked |
-11.1 | ||
| PS54 | ARG424
ASN461 ASN461 GLU458 GLU458 GLU458 ARG424 |
Conventional Hydrogen Bond
Conventional Hydrogen Bond Conventional Hydrogen Bond Pi-Anion Pi-Anion Pi-Anion Pi-Alkyl |
-10.1 | ||
| PS62 | ARG421
ARG457 ASP141 LEU460 TYR35 ARG457 ARG424 ARG454 ARG457 ARG457 LEU460 ARG421 |
Conventional Hydrogen Bond
Carbon Hydrogen Bond Pi-Anion Alkyl Pi-Alkyl Pi-Alkyl Pi-Alkyl Pi-Alkyl Pi-Alkyl Pi-Alkyl Pi-Alkyl Pi-Alkyl |
-9.8 | ||
| PS66 | THR205
TRP128 TRP128 TYR121 TRP128 TRP128 TRP128 TRP128 TRP128 TRP128 TYR121 |
Conventional Hydrogen Bond
Pi-Sulfur Pi-Sulfur Pi-Pi Stacked Pi-Pi Stacked Pi-Pi Stacked Pi-Pi Stacked Pi-Pi Stacked Pi-Pi Stacked Pi-Pi Stacked Pi-Alkyl |
-10.0 |
CONCLUSION
Among the 16 phenothiazine derivatives, the ligands PS15 and PS13 displayed high docking scores with the enzymes COX-1 (PDB: 3KK6) (-9.492 kcal/mol) and monoamine oxidase (PDB ID: 2BXR) (PDB: 3LN1) (-8.572 kcal/mol), respectively. Also revealed were the pharmacophores properties that support their biological function. The binding and affinity between the monoamine oxidase (PDB ID: 2BXR) enzyme and phenothiazine, which are responsible for the antidepressant activity features, have thus been identified with the use of these in silico techniques. We docked phenothiazine analogues with monoamine oxidase (PDB ID: 2BXR) enzymes to determine their anti-depressant activity features. It was found that the values of all the compounds fell within the usual range, and Lipinski’s rule of five was not violated. This means that the chemicals are expected to have high bioavailability.
References
- Ben-Levy R., Hooper S., Wilson R., Paterson H. F., Marshall C. J.. (1998) Nuclear export of the stress-activated protein kinase p38 mediated by its substrate MAPKAP kinase-2. Current Biology 8: 1049-1057 https://doi.org/10.1016/s0960-9822(98)70442-7 | Google Scholar
- Boehm J. C., Smietana J. M., Sorenson M. E., Garigipati R. S., Gallagher T. F., Sheldrake P. L., Bradbeer J., Badger A. M., Laydon J. T., Lee J. C., Hillegass L. M., GriPSold D. E., Breton J. J., Chabot-Fletcher M. C., Adams J. L., et al. (1996) 1-Substituted 4-Aryl-5-pyridinylimidazoles: A new class of cytokine suppressive drugs with low 5-lipoxygenase and monoamine oxidase (PDB ID: 2BXR) enzyme: Inhibitory potency. Journal of Medicinal Chemistry 39: 3929-3937 https://doi.org/10.1021/jm960415o | Google Scholar
- Chan W., Almasieh M., Catrinescu M., Levin L. A.. (2017) Cobalamin-associated superoxide scavenging in neuronal cells is a potential mechanism for vitamin B12-deprivation optic neuropathy. American Journal of Pathology 188: 160-172 https://doi.org/10.1016/j.ajpath.2017.08.032 | Google Scholar
- Cuadrado A., Nebreda A. R.. (2010) Mechanisms and functions of p38 MAPK signalling. Biochemical Journal 429: 403-417 https://doi.org/10.1042/bj20100323 | Google Scholar
- Field M. S., Kamynina E., Chon J., Stover P. J.. (2018) Nuclear folate metabolism. Annual Review of Nutrition 38: 219-243 https://doi.org/10.1146/annurev-nutr-071714-034441 | Google Scholar
- Gröber U., Kisters K., Schmidt J.. (2013) Neuroenhancement with vitamin B12—Underestimated neurological significance. Nutrients 5: 5031-5045 https://doi.org/10.3390/nu5125031 | Google Scholar
- Hobbenaghi R., Javanbakht J., Hosseini E., Mohammadi S., Rajabian M., Moayeri P., Hassan M. A., et al. (2013) RETRACTED ARTICLE: Neuropathological and neuroprotective features of vitamin B12 on the dorsal spinal ganglion of rats after the experimental crush of sciatic nerve: An experimental study. Diagnostic Pathology 8: Article 123 https://doi.org/10.1186/1746-1596-8-123 | Google Scholar
- Horiuchi K., Amizuka N., Takeshita S., Takamatsu H., Katsuura M., Ozawa H., Toyama Y., Bonewald L. F., Kudo A., et al. (1999) Identification and characterization of a novel protein, periostin, with restricted expression to periosteum and periodontal ligament and increased expression by transforming growth factor β. Journal of Bone and Mineral Research 14: 1239-1249 https://doi.org/10.1359/jbmr.1999.14.7.1239 | Google Scholar
- Kumar S., Boehm J., Lee J. C.. (2003) p38 MAP kinases: Key signalling molecules as therapeutic targets for inflammatory diseases. Nature Reviews Drug Discovery 2: 717-726 https://doi.org/10.1038/nrd1177 | Google Scholar
- Lee J. C., Young P. R.. (1996) Role of CSBP/p38/RK stress response kinase in LPS and cytokine signaling mechanisms. Journal of Leukocyte Biology 59: 152-157 https://doi.org/10.1002/jlb.59.2.152 | Google Scholar
- Ono K., Han J.. (2000) The p38 signal transduction pathway: Activation and function. Cellular Signalling 12: 1-13 https://doi.org/10.1016/s0898-6568(99)00071-6 | Google Scholar
- Ramesh G.. (2014) Novel therapeutic targets in neurodepression and neuropathic pain. Depression and Cell Signaling. https://doi.org/10.14800/ics.111 | Google Scholar
- Rathod R., Khaire A., Kale A., Joshi S.. (2016) A combined supplementation of vitamin B12 and n-3 polyunsaturated fatty acids across two generations improves nerve growth factor and vascular endothelial growth factor levels in the rat hippocampus. Neuroscience 339: 376-384 https://doi.org/10.1016/j.neuroscience.2016.10.018 | Google Scholar
- Streit W. J., Mrak R. E., Griffin W. S. T.. (2004) Microglia and neurodepression: A pathological perspective. Journal of Neuroinflammation 1: Article 14 https://doi.org/10.1186/1742-2094-1-14 | Google Scholar

